Web-based server, with platform-specific and command-line driven
AUTOMUTE PROTEIN PORTABLE
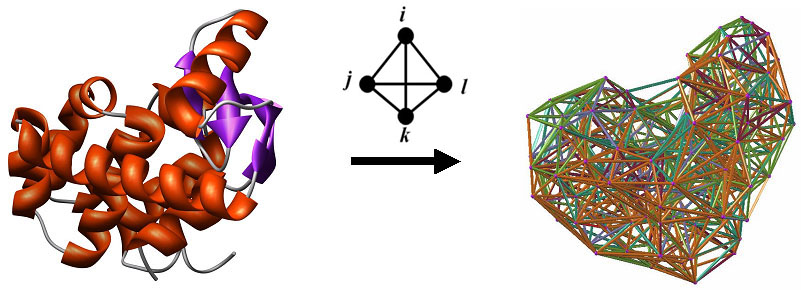
Here we introduce AUTO-MUTE 2.0, a portable alternative to the Statistical machine learning algorithms using the Java-based Weka Models were developed by implementing classification and regression Sequence similarity, were used to train the AUTO-MUTE predictors. Large sets of diverse mutations that have been studied experimentallyįor their functional effects, which occur in proteins that share low That include data derived from our in silico mutagenesis procedure. Single residue replacement can be represented as a vector of features For proteins with known 3D structures, any mutation defined by a The coarse graining of protein structures at the amino acid level. Implementing a computational mutagenesis technique that employs aįour-body, knowledge-based statistical potential function derived via Tools for predicting protein functional consequences upon mutation, by We previously developed the AUTO-MUTE server, an online set of Residue substitutions in proteins, and predictions of functional effectsĪre obtained via mathematical, rule-based, or statistical learning Each approach uniquely employsĮvolutionary, sequence, or structural information to characterize Well as reliable and immediate predictions for functional effects of Methods from the wet laboratory at a fraction of the time and cost, as Yielding efficient computational analogues to complement experimental Mutagenesis methodologies have been developed in recent years , Thermodynamic and physicochemical properties. Residues, to gain insights into mechanisms of protein folding, and toĪccumulate data needed for engineering new proteins with desired Or disease potential, to annotate structural or functional roles of
Opportunities to evaluate their effects on protein stability, activity, Site-directed mutagenesis experiments provide researchers with APA style: AUTO-MUTE 2.0: a portable framework with enhanced capabilities for predicting protein functional consequences upon mutation.AUTO-MUTE 2.0: a portable framework with enhanced capabilities for predicting protein functional consequences upon mutation." Retrieved from
AUTOMUTE PROTEIN FREE

In addition, we provide a perspective of how these methods will be beneficial for designing novel precision medicine approaches for several genetic disorders caused by mutations, such as cancer and neurodegenerative diseases. Here, we review these issues, highlighting new challenges required to improve current tools and to achieve more reliable predictions. Despite the large number of computational approaches for predicting the protein stability upon mutation, there are still critical unsolved problems: 1) the limited number of thermodynamic measurements for proteins provided by current databases 2) the large intrinsic variability of ΔΔG values due to different experimental conditions 3) biases in the development of predictive methods caused by ignoring the anti-symmetry of ΔΔG values between mutant and native protein forms 4) over-optimistic prediction performance, due to sequence similarity between proteins used in training and test datasets. Protein stability predictions are becoming essential in medicine to develop novel immunotherapeutic agents and for drug discovery.
